$6M gift powers new PKD clinical and research activities
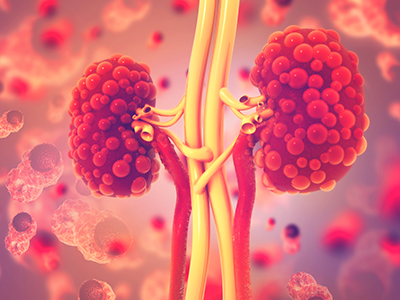
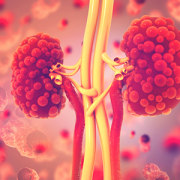
PKD is a genetic disorder characterized by clusters of fluid-filled sacs (cysts) multiplying and interfering with the kidneys’ ability to filter waste from the blood.
When Lisa M. Guay-Woodford, M.D., McGehee Joyce Professor of Pediatrics at Children’s National Hospital, considers a brand-new gift, she likens it to 6 million gallons of “rocket fuel” that will power new research to better understand polycystic kidney disease.
Dr. Guay-Woodford received a $5.7 million dollar gift to support PKD clinical and research activities. PKD is a genetic disorder characterized by clusters of fluid-filled sacs (cysts) multiplying and interfering with the kidneys’ ability to filter waste from the blood. The kidneys’ smooth surface transforms to a bumpy texture as the essential organs grow oversized and riddled with cysts.
The extraordinary generosity got its start in an ordinary clinical visit.
Dr. Guay-Woodford saw a young patient in her clinic at Children’s National a few times in 2015. The child’s diagnosis sparked a voyage of discovery for the patient’s extended family and, ultimately, they attended a presentation she gave during a regional meeting about PKD. That led to a telephone conversation and in-person meeting as they invited her to describe “the white space” between what was being done at the time to better understand PKD and what could be done.
“It’s the power of the art and science of medicine. They come to see people like me because of the science. If we can convey to patients and families that who they are and their unique concerns are really important to researchers, that becomes a powerful connection,” she says. “The art plus the science equals hope. That is what these families are looking for: We give people the latest insights about their disease because information is power.”
The infusion of new funding will strengthen the global initiative’s four pillars:
- Coordinated care for children and families impacted by renal cystic disease. The Inherited and Polycystic Kidney Disease (IPKD) program, launched September 2019, includes a cadre of experts working together as a team in the medical home so that “in a single, one-stop visit, Children’s National can address the myriad concerns they have,” she explains. A multi-disciplinary team that includes nephrologists, hepatologists and endocrinology experts meets weekly to ensure the Center of Excellence provides the highest-caliber patient care. The team includes genetic counselors to empower families with knowledge about genetic risks and testing opportunities. A nurse helps families navigate the maze of who to call about which issue. Psychologists help to ease anxiety. “There is stress. There is fear. There is pain that can be associated with this set of diseases. The good news is we can control their medical issues. The bad news is some children have difficulty coping. Our psychologists help children cope so they can be a child and do the normal things that children do,” she says.
- Strengthening global databases to capture PKD variations. The team will expand its outreach to other centers located around the world – including Australia, Europe, India and Latin America – caring for patients with both the recessive and dominant forms of polycystic kidney disease, to better understand the variety of ways the disease can manifest in children. “We really don’t know a lot about kids with the dominant form of the disease. How hard should we push to control their blood pressure, knowing that could ease symptoms? What are the ramifications of experiencing acute pain compared with chronic pain? How much do these pain flareups interfere with daily life and a child’s sense of self,” she asks. Capturing the nuances of the worldwide experience offers the power of harnessing even more data. And ensuring that teams collect data in a consistent way means each group would have the potential to extract the most useful information from database queries.
- Filling a ‘desperate need’ for biomarkers. Developing clinical trials for new therapies requires having biomarkers that indicate the disease course. Such biomarkers have been instrumental in personalizing care for patients with other chronic conditions. “We are in desperate need for such biomarkers, and this new funding will underwrite pilot studies to identify and validate these disease markers. The first bite at the apple will leverage our imaging data to identify promising biomarkers,” she says.
- Genetic mechanisms that trigger kidney disease. About 500,000 people in the U.S. have PKD. In many cases, children inherit a genetic mutation but, often, their genetic mutation develops spontaneously. Dr. Guay-Woodford’s research about the mechanisms that make certain inherited renal disorders lethal, such as autosomal recessive polycystic kidney disease, is recognized around the world. The fourth pillar of the new project provides funding to continue her lab’s research efforts to improve the mechanistic understanding of what triggers PKD.