Once overlooked cellular messengers could combat antibiotic resistance
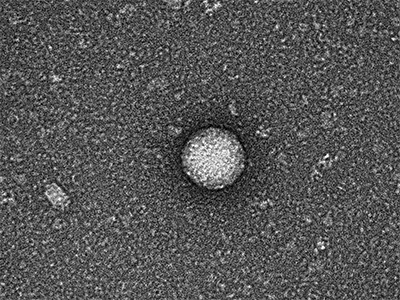
Children’s National Hospital researchers for the first time have isolated bacterial extracellular vesicles from the blood of healthy donors. The team theorizes that the solar eclipse lookalikes contain important signaling proteins and chromatin, DNA from the human host.
Children’s National Hospital researchers for the first time have isolated bacterial extracellular vesicles from the blood of healthy donors, a critical step to better understanding the way gut bacteria communicate with the rest of the body via the bloodstream.
For decades, researchers considered circulating bacterial extracellular vesicles as bothersome flotsam to be jettisoned as they sought to tease out how bacteria that reside in the gut whisper messages to the brain.
There is a growing appreciation that extracellular vesicles – particles that cells naturally release – actually facilitate intracellular communication.
“In the past, we thought they were garbage or noise,” says Robert J. Freishtat, M.D., MPH, associate director, Center for Genetic Medicine Research at Children’s National Research Institute. “It turns out what we throw away is not trash.”
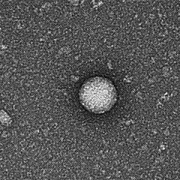
Kylie Krohmaly, a graduate student in Dr. Freishtat’s laboratory, has isolated from blood, extracellular vesicles from Escherichia coli and Haemophilus influenzae, common bacteria that colonize the gut, and validated the results via electron microscopy.
“The images are interesting because they look like they have a bit of a halo around them or penumbra,” Krohmaly says.
The team theorizes that the solar eclipse lookalikes contain important signaling proteins and chromatin, DNA from the human host.
“It’s the first time anyone has pulled them out of blood. Detecting them is one thing. Pulling them out is a critical step to understanding the language the microbiome uses as it speaks with its human host,” Dr. Freishtat adds.
Krohmaly’s technique is so promising that the Children’s National team filed a provisional patent.
The Children’s research team has devised a way to gum up the cellular works so that bacteria no longer become antibiotic resistant. Targeted bacteria retain the ability to make antibiotic-resistance RNA, but like a relay runner dropping rather than passing a baton, the bacteria are thwarted from advancing beyond that step. And, because that gene is turned off, the bacteria are newly sensitive to antibiotics – instead of resistant bacteria multiplying like clockwork these bacteria get killed.
“Our plan is to hijack this process in order to turn off antibiotic-resistance genes in bacteria,” Dr. Freishtat says. “Ultimately, if a child who has an ear infection can no longer take amoxicillin, the antibiotic would be given in tandem with the bacteria-derived booster to turn off bacteria’s ability to become antibiotic resistant. This one-two punch could become a novel way of addressing the antibiotic resistance process.”
ISEV2020 Annual Meeting presentation
(Timing may be subject to change due to COVID-19 safety precautions)
Oral with poster session 3: Neurological & ID
Saturday May 23, 2020, 5 p.m. to 5:05 p.m. (ET)
“Detection of bacterial extracellular vesicles in blood from healthy volunteers”
Kylie Krohmaly, lead author; Claire Hoptay, co-author; Andrea Hahn, M.D., MS, infectious disease specialist and co-author; Robert J. Freishtat, M.D., MPH, associate director, Center for Genetic Medicine Research at Children’s National Research Institute and senior author.