Prenatal COVID exposure associated with changes in newborn brain
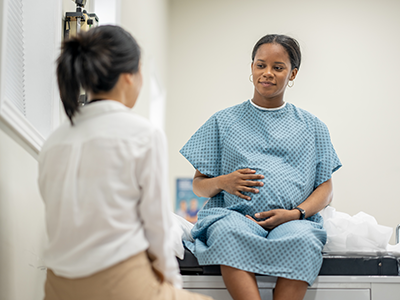
The team found differences in the brains of both infants whose mothers were infected with COVID while pregnant, as well as those born to mothers who did not test positive for the virus.
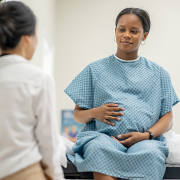
Babies born during the COVID-19 pandemic have differences in the size of certain structures in the brain, compared to infants born before the pandemic, according to a new study led by researchers at Children’s National Hospital.
The team found differences in the brains of both infants whose mothers were infected with COVID while pregnant, as well as those born to mothers who did not test positive for the virus, according to the study published in Cerebral Cortex.
The findings suggest that exposure to the coronavirus and being pregnant during the pandemic could play a role in shaping infant brain development, said Nickie Andescavage, M.D., the first author of the paper and associate chief for the Developing Brain Institute at Children’s National.
The fine print
The study’s authors looked at three groups of infants: 108 born before the pandemic; 47 exposed to COVID before birth; and 55 unexposed infants. In all cases, researchers performed magnetic resonance imaging (MRI) scans of the newborns’ brains during the first few weeks of life. The MRI scans, which are non-invasive and do not expose patients to radiation, provided 3D images of the brain, allowing doctors to calculate the volume of different areas.
Researchers found several differences in the brains of babies exposed to COVID. They had larger volumes of the gray matter that makes up the brain’s outermost layer, compared to the two other groups. In contrast, an inner area of the brain, known as deep gray matter, was smaller than in unexposed babies. These are areas that contain large numbers of neurons that generate and process signals throughout the brain. “Their brains formed differently if they were exposed to COVID,” said Dr. Andescavage, adding that “those exposed to COVID had unique signatures” in the brain.
Doctors also measured the depths of the folds in the babies’ brains – a way to determine how the brain is maturing during early development. Babies born to mothers who had COVID in pregnancy had deeper grooves in the frontal lobe, while babies born during the pandemic – even without being exposed to COVID – had increased folds and grooves throughout the brain, compared to babies born before the pandemic. “There was something about being born during the pandemic that changed how the brain developed,” Dr. Andescavage said.
What’s ahead
The study authors can’t fully explain what caused the differences in brain development in these babies, Dr. Andescavage said. But other studies have linked maternal stress and depression to changes in the newborn brain. In a future study, Dr. Andescavage and her colleagues will examine the relationship between infant brain development and how stress and anxiety during the pandemic may have played a role in early development.
Because the babies in the study were just a few weeks old, researchers don’t know if their altered brain development will affect how they learn or behave. Researchers plan to follow the children until age 6, allowing them to observe whether pandemic-era babies hit key developmental milestones on time, such as walking, talking, holding a crayon and learning the alphabet.
Researchers have been worried about the effect of COVID on the fetus since the beginning of the pandemic. Studies show that babies exposed to COVID in the womb may experience developmental impacts, and research is underway to better understand long-term outcomes.
Although the coronavirus rarely crosses the placenta to infect the fetus directly, there are other ways maternal infection can influence the developing baby. Dr. Andescavage said inflammation is one potential harm to a developing baby. In addition, if a pregnant woman becomes so sick that the levels of oxygen in her blood fall significantly, that can deprive the fetus of oxygen, she added.
In recent decades, studies of large populations have found that maternal infections with influenza and other viruses increased the risk of serious problems in children even years later, including autism, attention deficit hyperactivity disorder and schizophrenia, although the reasons behind the association are not well understood. Technology may allow doctors to answer a number of questions about COVID and the infant brain.
“With advanced imaging and MRI, we’re in a position now to be able to understand how the babies are developing in ways we never previously could,” Dr. Andescavage said. “That will better allow us to identify the exposures that may be harmful, and at what times babies may be especially vulnerable, to better position us to promote maternal wellness. This, in turn, helps infant wellness.”