Facial analysis technology successful in identifying Williams-Beuren syndrome in diverse populations

Image Credit: Darryl Leja, NHGRI.
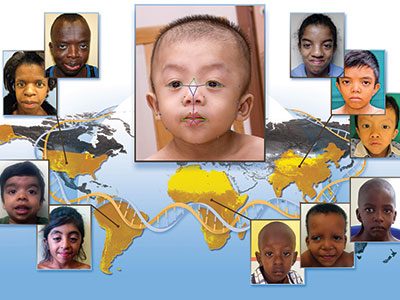
In an international study led by the National Human Genome Research Institute (NHGRI), researchers have successfully identified Williams-Beuren syndrome in diverse populations using clinical information and objective facial analysis technology developed by the Sheikh Zayed Institute for Pediatric Surgical Innovation at Children’s National.
The technology, which was featured by STAT as an ‘Editor’s Pick’ finalist in their recent competition to find the best innovation in science and medicine, enables users to compare the most relevant facial features characteristic of Williams-Beuren syndrome in diverse populations.
Williams-Beuren syndrome affects an estimated 1 in 7,500 to 10,000 people, with the most significant medical problems being cardiovascular, including high blood pressure. Though the syndrome is a genetic condition, most cases are not inherited. Signs and symptoms include intellectual disability and distinctive facial features including puffiness around the eyes, a short nose with a broad tip, full cheeks and a wide mouth with full lips.
Using the facial analysis technology, the researchers compared 286 African, Asian, Caucasian and Latin American children and adults with Williams-Beuren syndrome with 286 people of the same age, sex and ethnicity without the disease. They were able to correctly identify patients with the disease from each ethnic group with 95 percent or higher accuracy.
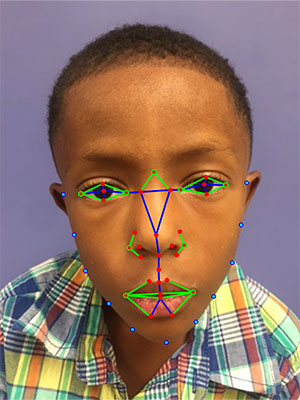
“Our algorithm found that the angle at the nose root is the most significant facial feature of the Williams-Beuren syndrome in all ethnic groups and also highlighted facial features that are relevant to diagnosing the syndrome in each group,” said Marius George Linguraru, D.Phil., developer of the facial analysis technology and an investigator in the study from Children’s National.
Linguraru and his team are working to create a simple tool that will enable doctors in clinics without state-of-the-art genetic facilities to take photos of their patients on a smartphone and receive instant results.
The technology was also highly accurate in identifying Noonan syndrome according to a study published in Sept. 2017, DiGeorge syndrome (22q11.2 deletion syndrome) in April 2017 and Down syndrome in Dec. 2016. The next study in the series will focus on Cornelia de Lange syndrome.