Researcher to decipher how viruses affect the developing brain with nearly $1M NIH award
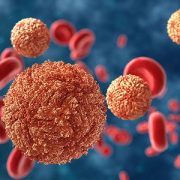
Zika virus in blood with red blood cells, a virus which causes Zika fever found in Brazil and other tropical countries.
The National Institutes of Health (NIH) awarded Children’s National Hospital nearly $1M of research support toward uncovering the specific cellular response that happens inside a developing brain once it is infected with a virus, including how the neuron gets infected, and how it dies or survives. The research is expected to gather critical information that can inform prenatal neuro-precision therapies to prevent or attenuate the effects of endemic and pandemic viruses in the future.
“We need to use all of the information we have from ongoing and past pandemics to prevent tomorrow’s public health crisis,” said Youssef Kousa, MS, D.O., Ph.D., neonatal critical care neurologist and physician-scientist at Children’s National. “There is still here a whole lot to learn and discover. We could eventually — and this is the vision that’s inspiring us — prevent neurodevelopmental disorders before a baby is born by understanding more about the interaction between the virus, mother, fetus, and environment, among other factors.”
Different viruses, including HIV, CMV, Zika and rubella, injure the developing brain in very similar ways. This line of work was fostered first by the clinical research team led by Adre du Plessis, M.B.Ch.B., and Sarah Mulkey, M.D., supported by Catherine Limperopoulos, Ph.D., chief and director of the Developing Brain Institute at Children’s National.
The clinical research findings then led to the NIH support, which then inspired more basic science research. Fast forward to now, Kousa will study how Zika affects the human brain and extrapolate what is learned and discovered for a broader understanding of neurovirology.
The research program is supported by senior scientists and advisors, including Tarik Haydar, Ph.D., and Eric Vilain, M.D., Ph.D., both at Children’s National and Avindra Nath, M.D., at NIH, as well as other leading researchers at various U.S. centers.
“This is a team effort;” added Kousa, “I’m thankful to have a group of pioneering and seasoned researchers engaged with me throughout this process to provide invaluable guidance.”
Many viruses can harm the developing brain when they replicate in the absence of host defenses, including the gene regulatory networks responsible for the neuronal response. As a result, viral infections can lead to brain injury and neurodevelopmental delays and disorders such as intellectual disability, seizures that are difficult to treat, and vision or hearing loss.
The big picture

Youssef Kousa, MS, D.O., Ph.D., neonatal critical care neurologist and physician-scientist at Children’s National.
The translational research supported by NIH with this award synergistically complements nationally recognized clinical research programs and ongoing prospective cohort studies at Children’s National to identify the full spectrum of neurodevelopmental clinical outcomes after prenatal Zika and other viral infections led by Dr. Mulkey and Roberta DeBiasi, M.D., M.S..
The award also builds upon strengths at the Children’s National Research & Innovation Campus, which is in proximity to federal science agencies. Children’s National experts from the Center for Genetic Medicine Research, known for pediatric genomic and precision medicine, joined forces with the Center of Neuroscience Research and the NIH-NINDS intramural research program to focus on examining prenatal and childhood neurological disorders.
Kousa received this competitive career development award from the National Institute of Neurological Disorders and Stroke of the National Institutes of Health under Award Number K08NS119882. The research content is solely the responsibility of the authors and does not necessarily represent the official views of the National Institutes of Health.
The hold-up in the field
Many neurodevelopmental disorders are caused by endemic viruses, such as CMV, and by viral pandemics, including rubella as seen in the 1960s and Zika since 2015. By studying Zika and other prenatal viral infections, Kousa and team hope to gain deeper biological understanding of the viral effects toward developing therapies for anticipating, treating and preventing virally induced prenatal brain injury in the long-term future.
To date, little is known about how viruses affect developing neurons and, as a result, prenatal brain injury is not yet treatable. To bridge the gap towards prenatal neuro-precision therapies, the research explores how genes regulate neuronal developmental and viral clearance by innovatively integrating three systems:
- Cerebral organoids, which illuminate how a neuron reacts to a viral infection
- Pre-clinical models that link prenatal brain injury to postnatal neurodevelopmental outcomes
- Populational genomics to identify human genetic risk or protective factors for prenatal brain injury
Given the scope and complexity of this issue, the international Zika Genetics Consortium, founded in 2015 by Kousa and a team of leading investigators across the world, provides critical samples and resources for the third arm of the research by performing comprehensive genomic analyses using sequencing data collected from diverse human populations throughout Central and South America, which are not as heavily sequenced as Western populations. Through partnerships with the Centers for Disease Control and Prevention and NIH, the consortium’s database and biorepository houses thousands of patient records and biospecimens for research studies to better understand how viruses affect the developing human brain.
“It is inspiring to imagine that, in the longer term, we could recognize early on the level of brain-injury risk faced by a developing fetus and have the tools to mitigate ensuing complications,” said Kousa. “What is driving this research is the vision that one day, brain injury could be prevented from happening before a baby is born.”