Understanding gut bacteria: forces for good (and sometimes evil)
In a paper published Sept. 11, 2019, in PLOS ONE, a multi-institutional research team led by George Washington University (GW) faculty found 157 different types of organisms (eight phyla, 18 classes, 23 orders, 38 families, 59 genera and 109 species) living inside the guts of healthy volunteers.
Back in 2015, an interdisciplinary group of research scientists made their case during a business pitch competition: They want to create a subscription-based service, much like 23andMe, through which people could send in samples for detailed analyses. The researchers would crunch that big data fast, using a speedy algorithm, and would send the consumer a detailed report.
But rather than ancestry testing via cheek swab, the team sought to determine the plethora of diverse bacterial species that reside inside an individual’s gut in their ultimate aim to improve public health.
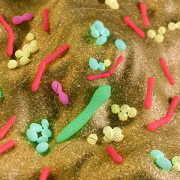
Hiroki Morizono, Ph.D., a member of that team, contributed detailed knowledge of Bacteroides, a key organism amid the diverse array of bacterial species that co-exist with humans, living inside our guts. These symbiotic bacteria convert the food we eat into elements that ensure their well-being as well as ours.
“Trillions of bacteria live in the gut. Bacteroides is one of the major bacterial species,” says Morizono, a principal investigator in the Center for Genetic Medicine Research at Children’s National in Washington, D.C. “In our guts they are usually good citizens. But if they enter our bloodstream, they turn evil; they’re in the wrong place. If you have a bacteroides infection, the mortality rate is 19%, and they resist most antibiotic treatments.”
The starting point for their project – as well as step one for better characterizing the relationship between gut bacteria and human disease – is taking an accurate census count of bacteria residing there.
In a paper published Sept. 11, 2019, in PLOS ONE, a multi-institutional research team led by George Washington University (GW) faculty did just that, finding 157 different types of organisms (eight phyla, 18 classes, 23 orders, 38 families, 59 genera and 109 species) living inside the guts of healthy volunteers.
The study participants were recruited through flyers on the GW Foggy Bottom campus and via emails. They jotted down what they ate and drank daily, including the brand, type and portion size. They complemented that food journal by providing fecal samples from which DNA was extracted. Fifty fecal metagenomics samples randomly selected from the Human Microbiome Project Phase I research were used for comparison purposes.
“The gut microbiome inherently is really, really cool. In the process of gathering this data, we are building a knowledge base. In this paper, we’re saying that by looking at healthy people, we should be able to establish a baseline about what a normal, healthy gut microbiome should look like and how things may change under different conditions,” Morizono adds.
And they picked a really, really cool name for their bacteria abundance profile: GutFeelingKB.
“KB is knowledge base. Our idea, it’s a gut feeling. It’s a bad joke,” he admits. “Drosophila researchers have the best names for their genes. No other biology group can compete. We, at least, tried.”
Next, the team will continue to collect samples to build out their bacteria baseline, associate it with clinical data, and then will start looking at the health implications for patients.
“One thing we could use this for is to understand how the bacterial population in the gut changes after antibiotic treatment. It’s like watching a forest regrow after a massive fire,” he says. “With probiotics, can we do things to encourage the right bacteria to grow?”
In addition to Morizono, study co-authors include Lead Author Charles H. King, and co-authors Hiral Desai, Allison C. Sylvetsky, Jonathan LoTempio, Shant Ayanyan, Jill Carrie, Keith A. Crandall, Brian C. Fochtman, Lusine Gasparyan, Naila Gulzar, Najy Issa, Lopa Mishra, Shuyun Rao, Yao Ren, Vahan Simonyan, Krista Smith and Senior Author, Raja Mazumder, all of George Washington University; Paul Howell and Sharanjit VedBrat, of KamTek Inc.; Konstantinos Krampis, of City University of New York; Joseph R. Pisegna, of VA Greater Los Angeles Healthcare System; and Michael D. Yao, of Washington DC VA Medical Center.
Financial support for research described in this post was provided by the National Science Foundation under award number 1546491 and the National Institutes of Health National Center for Advancing Translational Sciences under award number UL1TR000075.